Overview
This project demonstrates an end-to-end RNA-seq differential expression (DE) workflow in Python using pydeseq2 (DESeq2-like negative binomial modeling). The notebook covers:
- loading and validating a raw count matrix
- constructing metadata from GEO annotations (Series Matrix)
- running DE with an explicit contrast
- QC/visualization (PCA, clustered heatmap)
-
exporting results for downstream interpretation
- Links: GitHub
- Status: Complete
Why this project
This project asks which human genes change expression in SARS‑CoV‑2 positive vs negative samples (GSE152075) using a proper RNA‑seq DE workflow with pyDESeq2, producing interpretable QC and results visuals
Highlights
- Differential expression (pydeseq2)
- Count-based modeling using a negative binomial GLM
- Multiple testing correction (FDR / adjusted p-values)
- Results table extraction via DeseqStats(…).results_df
- QC and visualization
- PCA to inspect global sample structure (and detect technical effects)
- Heatmap / clustermap of top DE genes (pattern-level interpretation)
Result
| Figure |
|---|
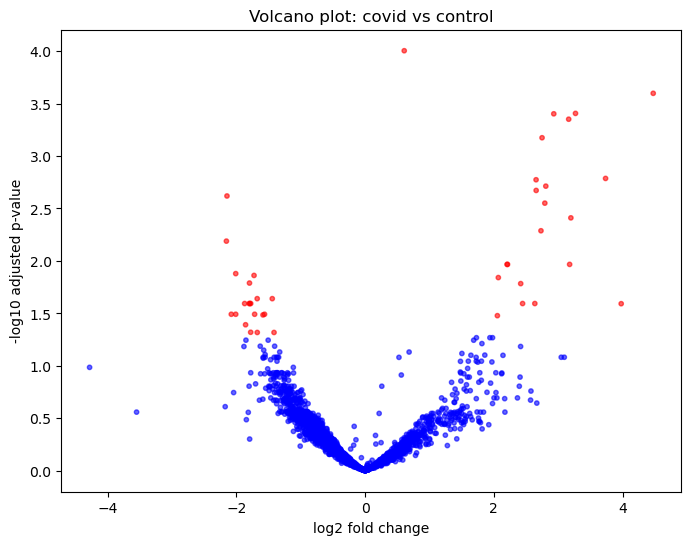 Figure 1: Volcano plot Tumor vs Normal |
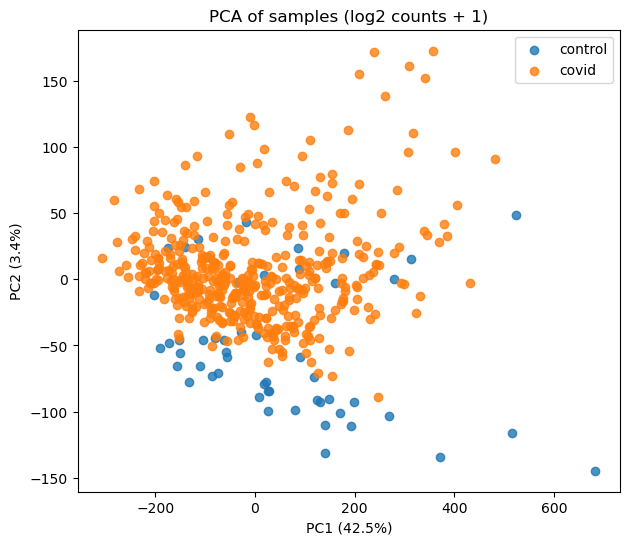 Figure 2: PCA plot Tumor vs Normal |
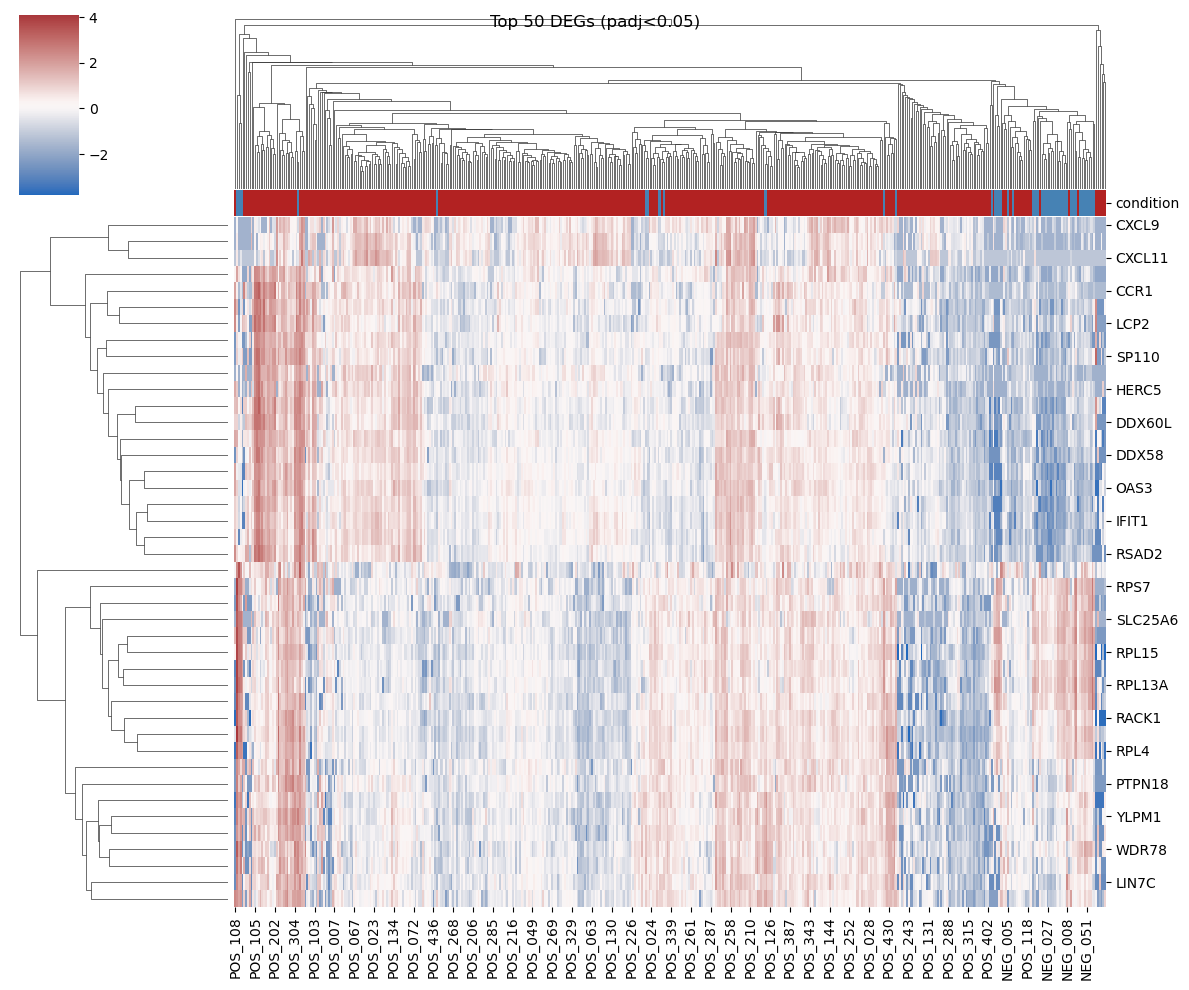 Figure 3: Top 20 genes based on p-value |
How to run
git clone https://github.com/Anwesha19-prog/RNA-seq-Differential-Expression
cd RNA-seq-Differential-Expression
pip install pandas numpy matplotlib seaborn scikit-learn pydeseq2
jupyter notebook Project-2_clean.ipynb