Portfolio
Introduction
Computational biology + translational oncology — building reproducible analyses and predictive models from clinical and multi-omics data.
What I do
- Modeling: survival / risk prediction, evaluation, interpretability
- Data & pipelines: omics preprocessing, QC, reproducible workflows
- ML for biology: prediction, validation, and interpretability (clear metrics, no leakage)
- Translation: turning research questions into analyses that support decisions
Featured projects
Project A — Multi omics integration (TCGA-BRCA)
This project integrates The Cancer Genome Atlas (TCGA) Breast Cancer (BRCA) multi-omics profiles—gene expression (RNA-seq), DNA methylation (450K), and copy-number variation (CNV) Read more →
Selected publication
Digital dynamic discrimination of primary colorectal cancer using systemic indocyanine green with near-infrared endoscopy (2021):
- Using real-time ICG near‑infrared endoscopy, the study shows that early fluorescence time‑curves (not single snapshots) reliably separate colorectal tumors from normal tissue and can even differentiate benign from malignant lesions, enabling a path toward software-assisted, intraoperative diagnosis.
Link
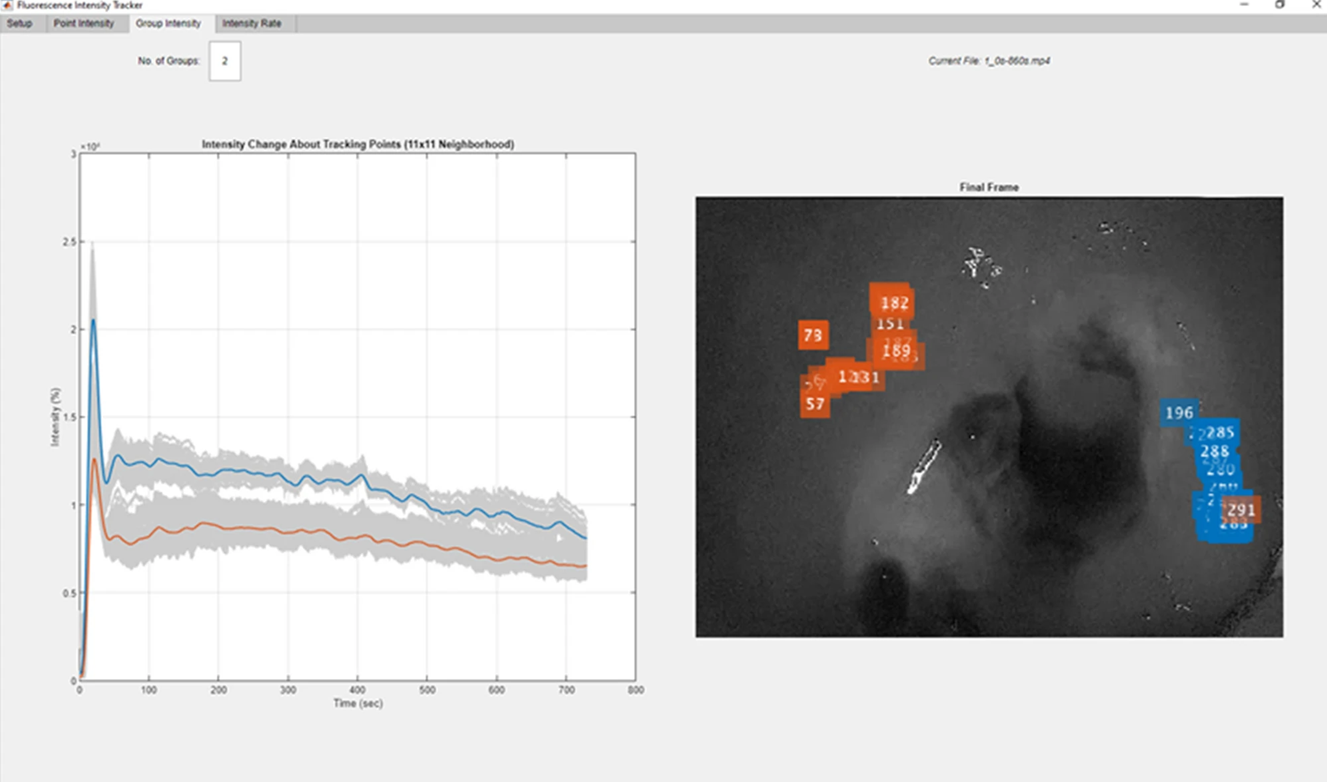 Figure 1: Screenshot of Fluorescence Tracker App (FTA, MATLAB®) showing post-hoc automated video-based classification of tumour (blue cluster) versus control tissue (orange cluster)
Figure 1: Screenshot of Fluorescence Tracker App (FTA, MATLAB®) showing post-hoc automated video-based classification of tumour (blue cluster) versus control tissue (orange cluster)
Currently
Open to computational biology / bioinformatics scientist roles (industry) and postdoc opportunities in translational computational biology.
