Overview
This repository contains a single, end-to-end Python notebook that integrates The Cancer Genome Atlas (TCGA) Breast Cancer (BRCA) multi-omics profiles—gene expression (RNA-seq), DNA methylation (450K), and copy-number variation (CNV) — to discover unsupervised patient groups and interpret them biologically.
- Links - Github
- Status: Complete
Why this project
Projects goals:
- Build a clean, reproducible multi-omics integration workflow in Python
- Identify molecularly distinct tumor groups using unsupervised learning
- Provide biologically grounded interpretation using canonical BRCA markers, CNV driver loci, and pathway enrichment
- Demonstrate practical skills in data harmonization, preprocessing, integration, clustering, and interpretation
Highlights
- Multi-omics data handling (RNA + methylation + CNV)
- Sample harmonization with TCGA barcodes
- Robust preprocessing and missing-value handling
- Dimensionality reduction (PCA) and integration (early integration of PCs)
- Unsupervised clustering with model selection (silhouette)
-
Biological interpretation via:
- canonical marker genes,
- CNV driver loci inspection,
- pathway enrichment (Hallmark)
Result
| Figure |
|---|
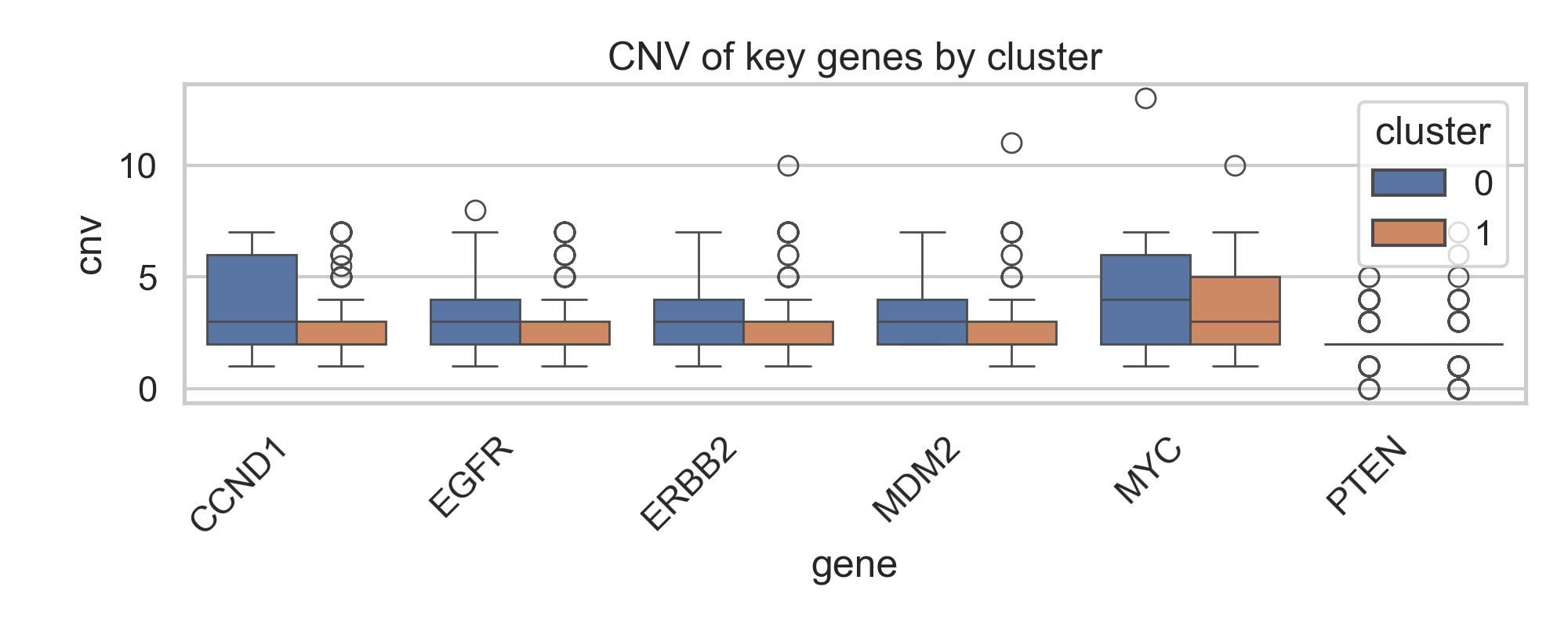 Figure 1: CNV of key genes by cluster |
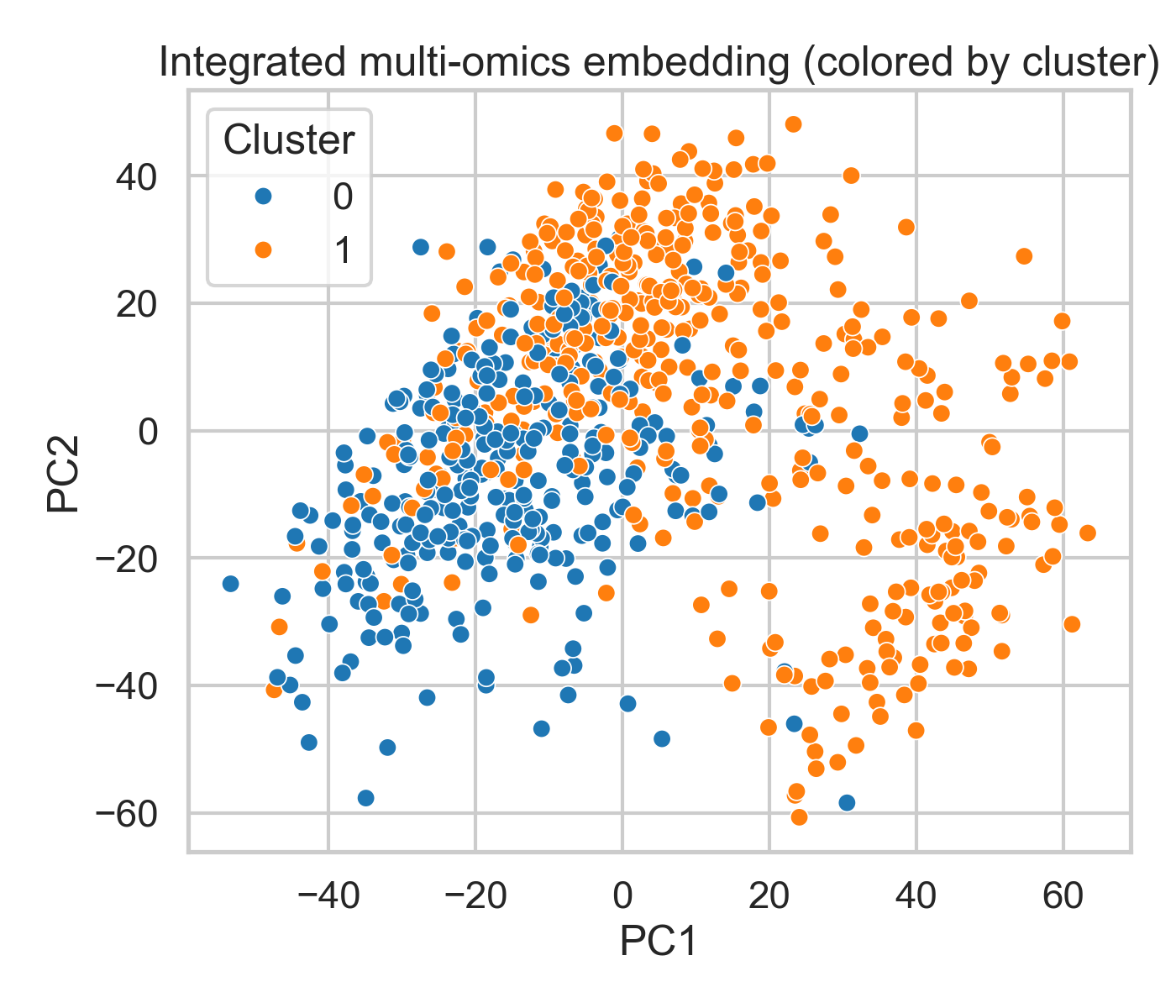 Figure 2: Integrated multi-omics embedding (colored by cluster) |
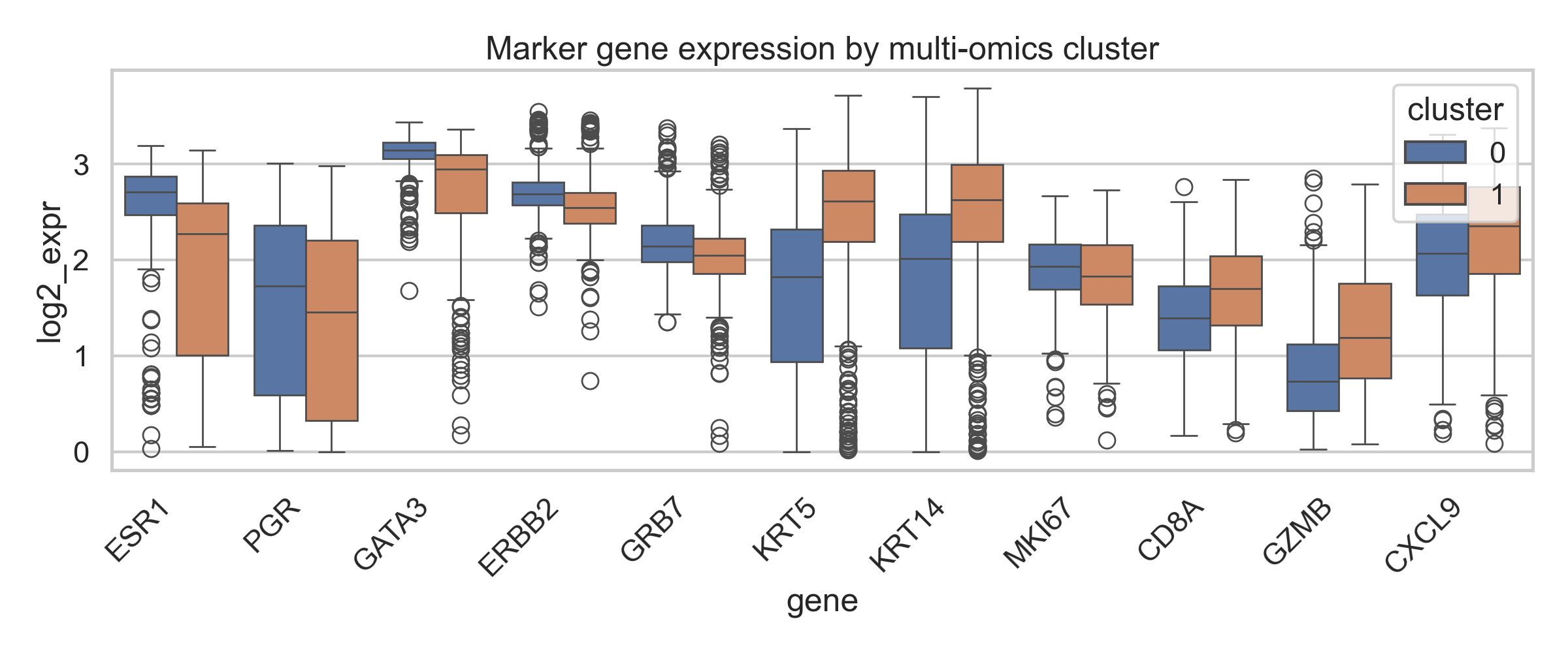 Figure 3: Marker gene expression by multi-omics cluster |
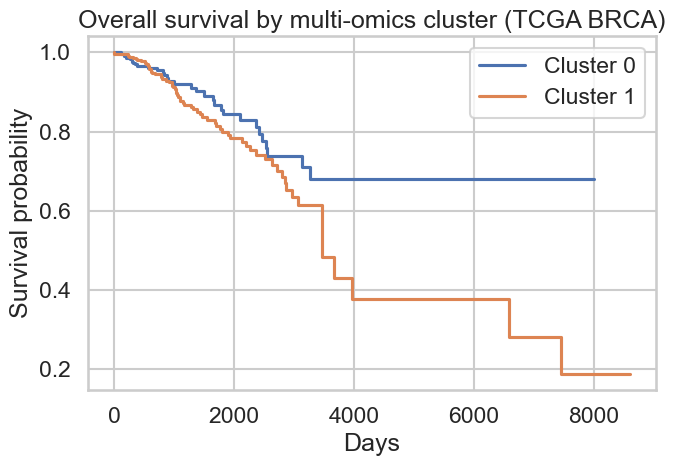 Figure 4: Overall survival by multi-omics cluster (TCGA BRCA) |
How to run
git clone https://github.com/Anwesha19-prog/TCGA-BRCA-Multi-Omics-Intergration
cd TCGA-BRCA-Multi-Omics-Intergration
python -m requirements.txt
jupyter notebook Project-3.ipynb
